Rolling hash, Rabin Karp, palindromes, rsync and others
Rolling hash is a neat idea found in the Rabin-Karp string matching algorithm which is easy to grasp and useful in quite a few different contexts.
As before, the most interesting part of the article is the problem section. Discuss them in the comment section, but first let's go through a few applications.
The Rabin-Karp algorithm
Given a string P of length n and a string S of length m find out all the occurrences of P within S
The algorithm works by building a fingerprint for each substring of S of length n. It checks if any of them match the fingerprint of P. Implementing this directly leads to a O(n*m) solution which isn't faster than the naive matching algorithm (in fact it may be slower).
The insight is that you can efficiently compute the hash value of a substring using the hash value of previous substring if you use the right hash function. A polynomial with the string characters as coefficients works well. Let ![H(c)=c[0]a^{n-1} + c[1]a^{n-2}+...+c[n-1] H(c)=c[0]a^{n-1} + c[1]a^{n-2}+...+c[n-1]](https://www.infoarena.ro/static/images/latex/13dd9ac43efaa22b2a9a43b253a955ae_3.5pt.gif) be the hash function for the string c.
be the hash function for the string c.
All the operations are done modulo a prime number so that we don’t have to deal with large integers.
Let’s see what happens when we go from H(S[i..i+n-1]) to H(S[i+1..i+n]):
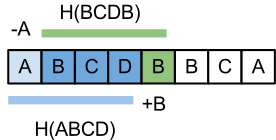
![H(S[i+1..i+n])= S[i+1]a^{n-1}+S[i+2]a^{n-2}+.. + S[i+n]
H(S[i+1..i+n])= S[i+1]a^{n-1}+S[i+2]a^{n-2}+.. + S[i+n]](https://www.infoarena.ro/static/images/latex/9786aaad72eea4d243845dabcfee26d1_3.5pt.gif)
Adding and substracting ![S[i]a^n S[i]a^n](https://www.infoarena.ro/static/images/latex/cda5ca2b5683782c5203ae8ba6ea2d5b_3.5pt.gif) we get
we get
![a(S[i]a^{n-1}+S[i+1]a^{n-2}+ S[i+2]a^{n-3}+ .. + S[i+n -1])+S[i+n]-S[i]a^{n} =
a(S[i]a^{n-1}+S[i+1]a^{n-2}+ S[i+2]a^{n-3}+ .. + S[i+n -1])+S[i+n]-S[i]a^{n} =](https://www.infoarena.ro/static/images/latex/9661b3d5405c73aaa4be4ebd8d839bd5_3.5pt.gif)
![aH(S[i..i+n-1])-S[i]a^{n}+S[i+n]
aH(S[i..i+n-1])-S[i]a^{n}+S[i+n]](https://www.infoarena.ro/static/images/latex/78f313b7a35a302ca083edc7b62fdd4b_3.5pt.gif)
Deleting a character and appending another one corresponds to adding a number and subtracting a number in our hashing algorithm. The complexity of the algorithm is O(n).
If S and P are strings of digits and a is 10, our problem maps to finding numbers in a string of digits.
Let's go through an example.
Let P = 53424, S = 3249753424234837 and a = 10
3249753424234837
32497
24975 = 10 * 32497 - 3 * 10000 + 5
49753 = 10 * 24975 - 2 * 10000 + 3
97534 = 10 * 49753 - 4 * 10000 + 4
...
53424 // match
...Here's some code:
an = 1
rolling_hash = 0
for i in range(0, n):
rolling_hash = (rolling_hash * a + S[i]) % MOD
an = (an * a) % MOD
if rolling_hash == hash_p:
print "match"
for i in range(1, m - n + 1):
rolling_hash = (rolling_hash * a + S[i + n - 1] - an * S[i - 1]) % MOD
if rolling_hash == hash_p:
print "match"Longest common substring
Given two strings A and B, compute their longest common substring.
Let's solve a simpler problem: Given two strings A and B, and a number X find if they have a common sequence of length X. We use the rolling hash technique to find the hash codes for all X length substrings of A and B. Then we check if there is any common hash code between the two sets, this means A and B share a sequence of length X. We then use binary search to find the largest X.
The complexity of this algorithm is O(n log n).
rsync
The linux utility is used to keep two copies of the same file in sync by copying around deltas. Instead of replicating the whole changed file, rsync segments the file into fixed size chunks, computes a rolling hash for each chunk. On the client side computes rolling hashes for all chunks of the target file including overlapping ones. Then it checks which chunks are already there, which chunks need to be deleted and which chunks need to be copied over. This improves the transfer speed as you only copy over deltas and some chunk fingerprints. Here's a better description from wikipedia.
Largest palindrome
Given a string of length n, find out its largest palindromic substring.
The naive solution is O(n3). For every possible substring it tests in linear time if it's a palindrome.
A smarter solution tries every position in the original string as the center of a palindrome and extends to the left and to the right as long as the corresponding characters match. This solution takes O(n2) time.
There's a O(n log n) solution that involves rolling hash. Given X we can test if there is any substring S' of S of length 2X such that H(S') == H(reverse(S')). The identity means that S contains a palindromic sequence of length at least 2X. What is left is to find out the maximum such X. We can use binary search to do that efficiently. The overall complexity is O(n log n).
Substrings of variable length
The polynomial function can be used to compute hashes for substrings of varying length
Let’s choose ![H(S[i..j]) = S[i]+S[i+1]a+S[i+2]a{2}+...+S[j]a^{j-i} H(S[i..j]) = S[i]+S[i+1]a+S[i+2]a{2}+...+S[j]a^{j-i}](https://www.infoarena.ro/static/images/latex/59151c9b034cbfc199631f11a4e03c83_3.5pt.gif) Then we compute the array h which contains cummulative hashes:
Then we compute the array h which contains cummulative hashes: ![h[k]=S[0]+S[1]a+S[2]a^{2}+...+S[k]a^{k} h[k]=S[0]+S[1]a+S[2]a^{2}+...+S[k]a^{k}](https://www.infoarena.ro/static/images/latex/c9dc20a1a9f14e076ec0c25d7985e913_3.5pt.gif) . It’s easy to see that
. It’s easy to see that ![H(S[i..j])=(h[j]-h[i-1])/a^i H(S[i..j])=(h[j]-h[i-1])/a^i](https://www.infoarena.ro/static/images/latex/e8a5105ec8fcf8b80150e575a526347e_3.5pt.gif)
After spending O(n) time on preprocessing h, we have an efficient way to compute the hash code for any substring of S.
The division operation modulo a prime needs some number theory insight. You can do it using Fermat's little theorem or using the Extended Euclidean Algorithm. If you need to process a lot of sub strings it's useful to pre process all the values for 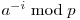 .
.
Caveats
Hashing collisions are a problem. One way to deal with them is to check for a match when two fingerprints are equal, but this makes the solution inefficient if there are lots of matches. Another way is to use more hashing functions to decrease the collision probability.
Alternatives
Suffix trees or suffix arrays are very good tools to tackle string matching problems. I prefer hashing as the code is usually simpler.
Additional problems
- Given a string of length n, count the number of palindromic substring of S. Solve better than O(n2).
- Given a string of length n, find the longest substring that is repeated at least k times. Solve in O(n log n)
- Given a bitmap, find out the largest square that’s repeated in the bitmap.
- Given a string S figure out if the string is periodic. (there is a p such that for any i we have that s[i] == s[i + p]) For example abcabcab is periodic with p = 3. O(n log n)
- Given two equal length strings, figure out if one is a rotation of the other. O(n)
- Given two polygons find out if they are similar polygons. O(n)
- Given a string, find it's longest periodic prefix. O(n log n) For aaabaaabcde the answer is aaabaaab
- Given a tree T with character labeled nodes and a given string P count how many downward paths match P. O(n)
